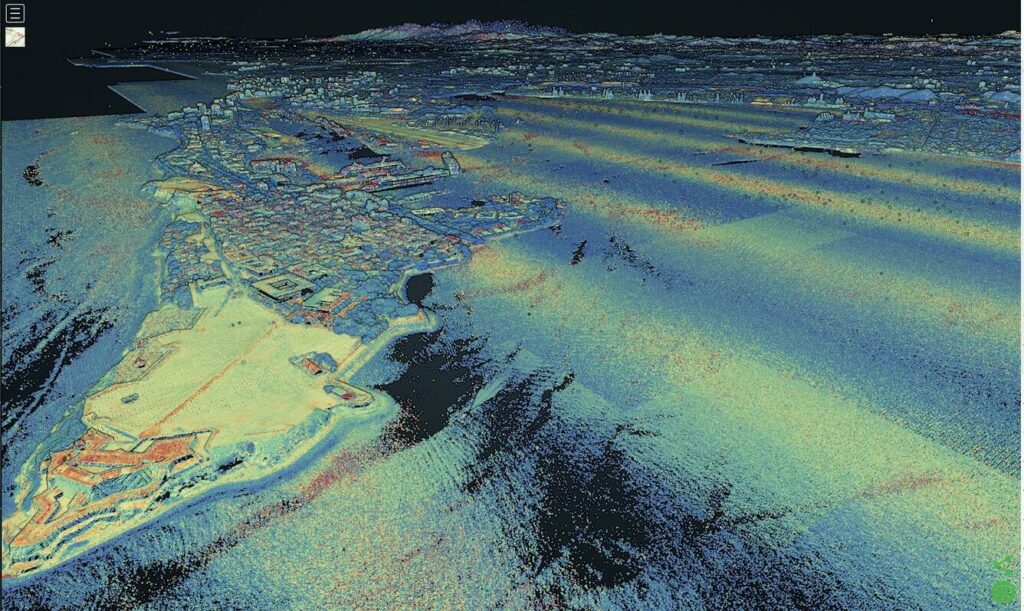
In part one I described some of the issues I had on a recent project that applied deep learning to geographic feature recognition in LiDAR and the file sizes of such data. This time I want to talk about training data, how important it is, and how little there is for this type of problem.
One of the most important things in deep learning is having both quality training data and a good amount of such data. I have actually written a previous post about the importance of quality data that you can read here. At a bare minimum, you should typically have around one thousand samples of each object class you want to train a model to recognize. Object classes in this case are geographic feature types that we want to recognize.
Training Data Characteristics
These samples should mirror the characteristics of the data that your model will come across during classification tasks. With LiDAR in GeoTIFF format, the training data should be similar in resolution (0.5 meters in this case) and bit depth (32-bit) to the area for testing. There should be variability in the training data. Convolutional neural networks are NOT rotational invariant, meaning that unless you train your model on samples at different angles, it will not automatically recognize features. In this case, your training LiDAR features should be rotated at different angles to account for differences in projection or north direction.
Balanced Numbers of Features per Class
Your training data should also be balanced, meaning that each class should have roughly the same number of training images where possible. You can perform some tasks that we will talk about in a bit to help with this, but generally if your model is unbalanced, it could mistakenly “lean” towards one class more than others.
Related to this, when generating your training data, you should make sure your training and testing data contain samples from each of your object classes. Especially when imbalanced, it is very easy to use something like train_test_split from the sklearn library and have it generate a training set that misses some object classes. You should also shuffle your training data so that the samples from each class are sufficiently randomized and the model does not assume that features will appear in a specific order. To alleviate this, make sure you pass in something like:
… = train_test_split(…, shuffle=True, stratify=labels)where labels is a list of your object classes. For most cases this will ensure the order of your samples is sufficiently randomized and that your training/testing data contains samples of each object class.
Complexity
Geographic features also have a varying complexity in how they appear in LiDAR. The same feature can “look” differently based on its size or other characteristics. Humans will always look like humans, so training on them is fairly simple. Geographic features can be in the same class but appear differently based on factors such as weathering, erosion, vegetation, and even if it’s a riverbed that is dry part of the year. This increased complexity means that samples need to have enough variability, even in the same object class, for proper training.
Lack of Training Data for This Project
With all of this out of the way, let us now talk about the issues that faced this particular project. First of all, there is not a lot of labeled training data for these types of features. At all. I used various search engines, ChatGPT, even bought a bucket of KFC so I could try to throw some bones to lead me to training data (although I don’t know voodoo so probably read them wrong).
There is a lot of data about these geographic features out there, but not labeled AND in LiDAR format. There is an abundance of photographs of these features. There are paper maps of these features. There are research papers with drawings of these features. I even found some GIS data that had polygons of these features, but the matching elevation model was too low of a resolution to be useful.
In the end I was only able to find a single dataset that matched the bit depth and spatial resolution that matched the test data. There were a couple of problems with this dataset though. First, it only had three feature classes out of a dozen or more. Second, the number of samples of each of these three classes were way imbalanced. It broke down like this:
- Class 1 – 123 samples
- Class 2 – 2,214 samples
- Class 3 – 9,700 samples
Realistically we should have just tried for Classes 2 and 3, but decided to try to use various techniques to help with the imbalance. Plus, since it was a bit of a research project, we felt it would be interesting to see what would happen.
Data Augmentation
There are a few different methods of data augmentation you can do to add more training samples, especially with raster data. Data augmentation is a technique where you generate new samples from existing data so that you can enhance your model’s generalization and generate more data for training. The key part of this is making sure what the methods you use do not change the object class of the training sample.
Geometric Transformations
The first thing you can do with raster data is to apply geometric transformations (again, as long as they do not change the object class of your training sample). Randomly rotating your training images can help with the rotational invariance mentioned above. You can also flip your images, change their scale, and even crop them as long as the feature in the training sample remains.
You can gain several benefits from applying geometric transformations to your training data. If your features can appear at different sizes, scaling transforms can help your model become better generalized on feature size. With LiDAR data, suppose someone did not generate the scene with North at the top. Here, random rotations can help the model generalize to rotation so it can better detect features regardless of angle.
Spatial Relationships
Regardless of what type of augmentations you apply to LiDAR, you have to be mindful that you do not change the spatial characteristics of the data. Consider color space augmentation, something that is common with other areas of deep learning and computer vision. With LiDAR, modifying the brightness/contrast would actually be changing the elevation and/or reflectance values of the data. In some applications, especially those highly based on reflectance values such as detecting types of vegetation, this might be useful. In high-resolution geography, you could end up altering the training data in such a way that it no longer represents real-world features.
Wrapping Up
I think I will end this one here as it got longer than I expected and I’m tired of typing 😉 Next time I’ll cover issues with image processing libraries and 32-bit LiDAR data.